Genomics platform to understand the varied types of disease progression of patients with COVID-19, which will help to identify what treatment is best suited to individual patients.
The ID Predict platform – led by University of Melbourne researchers at the Doherty Institute in collaboration with computational biologists at the Melbourne School of Engineering and the Water & Eliza Hall Institute, and international genomics company Illumina – will combine cutting-edge human and microbial genomics with multi-disciplinary technology advances to create a platform that is essential to the COVID-19 response, and rapidly deployable for future infectious disease outbreaks.
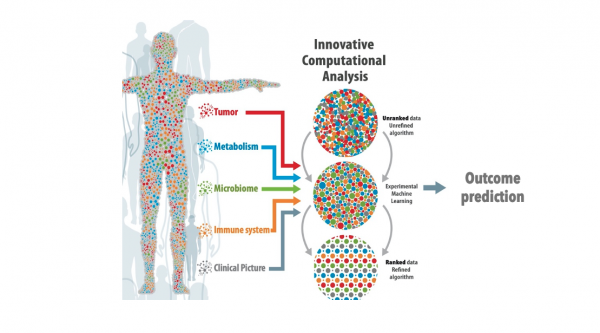
Public health genomics has provided insight on community transmission and spread, but understanding of why people respond to the virus so differently is still largely unknown. The ID Predict platform will examine aspects such as immune cell composition and function, virus parameters, microbiome diversity and metabolic activities, as well as co-morbidities to create deep biological and immunological profiling of patients. Innovative computational approaches will then be applied to these data sets to develop decision-making tools capable of predicting disease outcomes in individual patients and tailoring treatment options.
Rather than a ‘one-size-fits-all’ approach, this platform will allow physicians to personalise treatments to their patients. This will aid in ensuring that each patient is rapidly subjected to the best treatment, tailored to their own unique genetic make up.

Principal Investigator: Professor Sammy Bedoui
The Bedoui Lab uses models of viral and bacterial infection to study how the innate and the adaptive immune system interact. Key foci are to understand how innate cells sense pathogens and how this information is integrated into protective adaptive T cell responses.